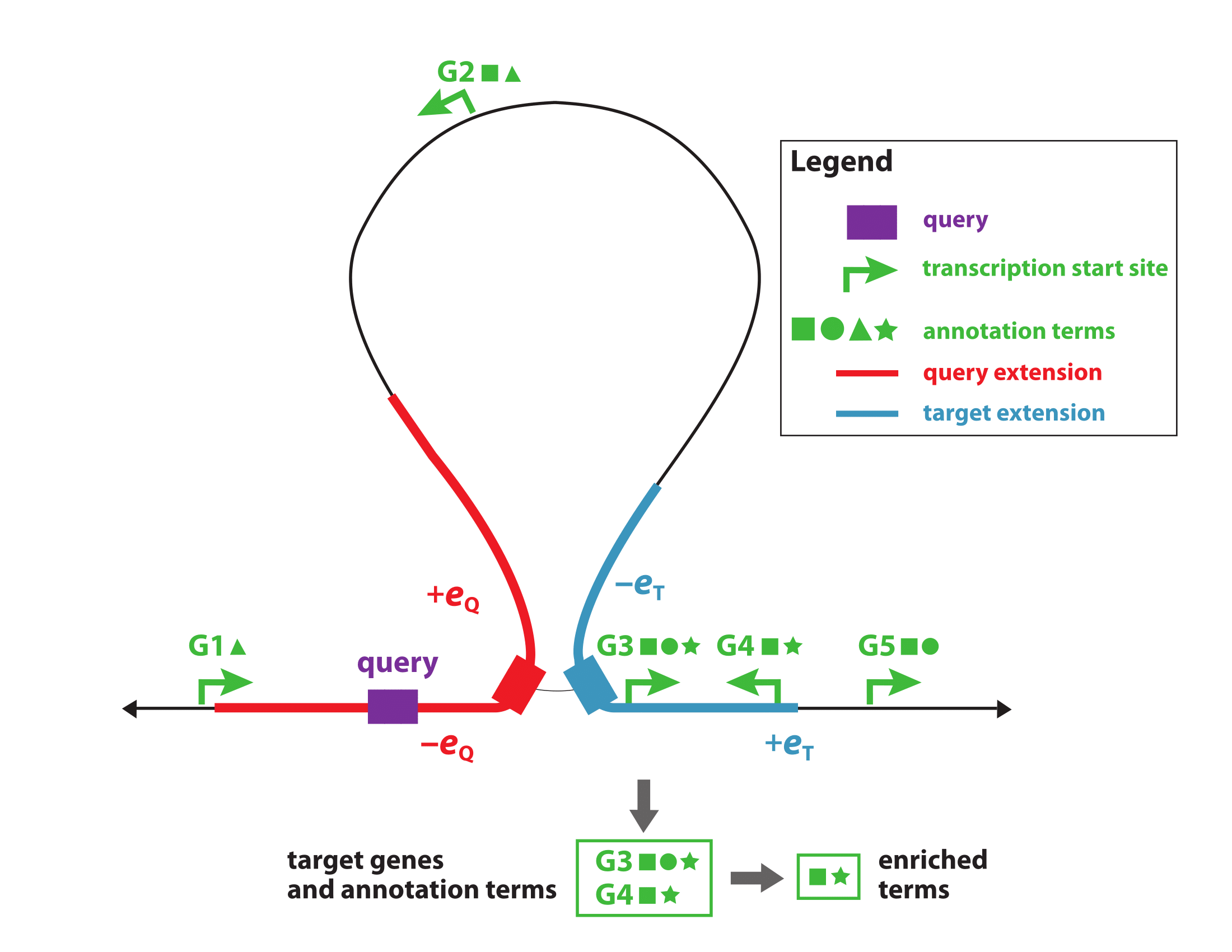
Chicco D, Bi HS, Reimand J, Hoffman MM. 2019. BEHST – Genomic set enrichment analysis enhanced through integration of chromatin long-range interactions. preprint.

The web application of BEHST is publically available on behst.hoffmanlab.org
BEHST can run on any Linux and Mac computers. You can find the installation instructions on the BEHST Bitbucket webpage.
You can find the source code repository on the BEHST Bitbucket webpage.
For support of BEHST, please write to the BEHST-users mailing list, rather than writing the authors directly. Using the mailing list will get your question answered more quickly. It also allows us to pool knowledge and reduce getting the same inquiries over and over. Questions sent to the mailing list will receive a higher priority than those sent to us individually.
If you do not want to read discussions about other people's use of BEHST, but would like to hear about new releases and other important information, please subscribe to the BEHST-announce mailing list mailing list.
BEHST was developed by Davide Chicco, Haixin Sarah Bi, Juri Reimand, and Michael M. Hoffman at the Hoffman Lab of the Princess Margaret Cancer Centre (Toronto, Ontario, Canada).